Software to align dna sequences9/17/2023  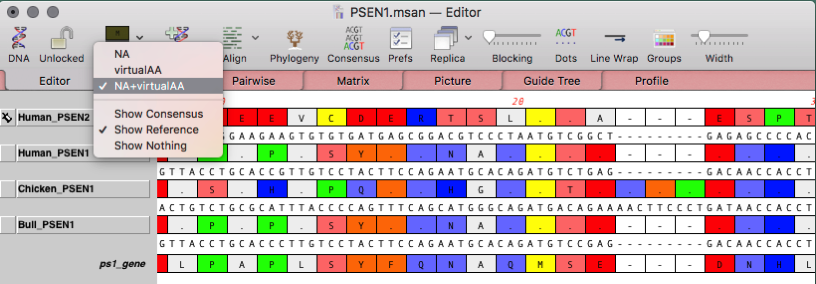
This will help us understand the concept of sequence alignment and how to program it using Biopython. Let us write an example to find the sequence alignment of two simple and hypothetical sequences using pairwise module. Biopython applies the best algorithm to find the alignment sequence and it is par with other software. Pairwise is easy to understand and exceptional to infer from the resulting sequence alignment.īiopython provides a special module, Bio.pairwise2 to identify the alignment sequence using pairwise method. Pairwise sequence alignment compares only two sequences at a time and provides best possible sequence alignments. Here, parse method returns iterable alignment object and it can be iterated to get actual alignments. > alignments = AlignIO.parse(open("PF18225_seed.txt"), "stockholm") If the input sequence alignment format contains more than one sequence alignment, then we need to use parse method instead of read method as specified below − In multiple sequence alignment concept, two or more sequences are compared for best subsequence matches between them and results in multiple sequence alignment in a single file. In general, most of the sequence alignment files contain single alignment data and it is enough to use read method to parse it. TRRHGPESFRFWLERQPVEARDRIYAIDRSGAEILDRPIPRGMAPLFKVLSFRNREDQGLVNNKPĪINRNTQQLTQDLRAMPNWSLRFVYIVDRNNQDLLKRPLPPGIMVLAPRLTAKHPYDKVQDRNRKĪVNATEREFTERIRTLPHWARRNVFVLDSQGFEIFDRELPSPVADLMRKLDLDRPFKKLERKNRT We can also check the sequences (SeqRecord) available in the alignment as well as below − TRRHGPESFRFWLERQPVEARDRIYAIDRSGAEILDRPIPRGMA.NKP A0A0X3UC67_9GAMM/57-121ĪINRNTQQLTQDLRAMPNWSLRFVYIVDRNNQDLLKRPLPPGIM.NRK B3PFT7_CELJU/62-126ĪVNATEREFTERIRTLPHWARRNVFVLDSQGFEIFDRELPSPVA.NRT K4KEM7_SIMAS/61-125 MQNTPAERLPAIIEKAKSKHDINVWLLDRQGRDLLEQRVPAKVA.EGP B7RZ31_9GAMM/59-123ĪKQRGIAGLEEWLHRLDHSEAIPIFLIDEAGKDLLEREVPADIT.KKP A0A0C3NPG9_9PROT/58-119ĪRRHGQEYFQQWLERQPKKVKEQVFAVDQFGRELLGRPLPEDMA.KKP A0A143H元7_9GAMM/57-121 SingleLetterAlphabet() alignment with 6 rows and 65 columns > alignment = AlignIO.read(open("PF18225_seed.txt"), "stockholm") parse method returns iterable alignment object similar to parse method in Bio.SeqIO module. If the given file contain many alignment, we can use parse method. 
read method is used to read single alignment data available in the given file. Let us try to read the downloaded sequence alignment file using Bio.AlignIO as below − Import Bio.AlignIO module Step 3 − Go to alignment section and download the sequence alignment file in Stockholm format (PF18225_seed.txt). Here, we have selected/clicked PF18225 and it opens go to and shows complete details about it, including sequence alignments. It contains minimal data and enables us to work easily with the alignment. Step 2 − Choose any one family having less number of seed value. It will show all the Pfam families in alphabetical order. 
Step 1 − Open your favorite browser and go to website. To download the sample file, follow the below steps − Bio.AlignIO provides API similar to Bio.SeqIO except that the Bio.SeqIO works on the sequence data and Bio.AlignIO works on the sequence alignment data.īefore starting to learn, let us download a sample sequence alignment file from the Internet. In bioinformatics, there are lot of formats available to specify the sequence alignment data similar to earlier learned sequence data. Let us learn some of the important features provided by Biopython in this chapter − Parsing Sequence Alignmentīiopython provides a module, Bio.AlignIO to read and write sequence alignments. Biopython provides extensive support for sequence alignment. Identifying the similar region enables us to infer a lot of information like what traits are conserved between species, how close different species genetically are, how species evolve, etc. Sequence alignment is the process of arranging two or more sequences (of DNA, RNA or protein sequences) in a specific order to identify the region of similarity between them. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
